Recombinant DNA
The problem is that those nucleotide sequences are embedded in an enormous genome.
The development of recombinant DNA technology provided a solution to this isolation problem.
Over the past thirty years this technology has matured into a powerful analytical tool which finds application in a diverse set of scientific disciplines ranging from ecology and evolution to mathematics and information science. It is this broad applicability that makes this technology so important to your undergraduate education.
Recombinant DNA technology originated with the collaboration between biochemists and geneticists to understand the Central Dogma. A favorite model system of the time was the prokaryote E. coli and its attendant bacteriophage. Among the phenomena being studied was that of
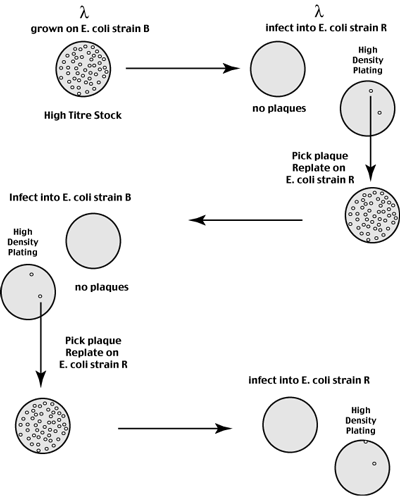
Bacteriophage lambda Host Restriction
a lambda phage stock grown on one strain of E. coli (strain B) infects that strain with high efficiency.
This same phage stock infects a second E. coli strain R with very low efficiency (1 in 203 plaques grow)
Recovery and re-titreing of progeny phage from one of these rare plaques shows that they infect the R strain with high efficiency, but infect the B strain with very low efficiency (1 in 203 plaques grow)
Recovery and re-titreing of progeny phage from one of these rare plaques shows the same phenomena described above.
Phage grown in one strain infect that strain with high efficiency but infect the other strain with low efficiency.